Projects
Selected work across gene regulation, multimodal learning, and network inference.
GRIDOT
Optimal Transport + Graph-based Granger Causality for GRN inference

GRIDOT
Optimal Transport + Graph-based Granger Causality for GRN inference
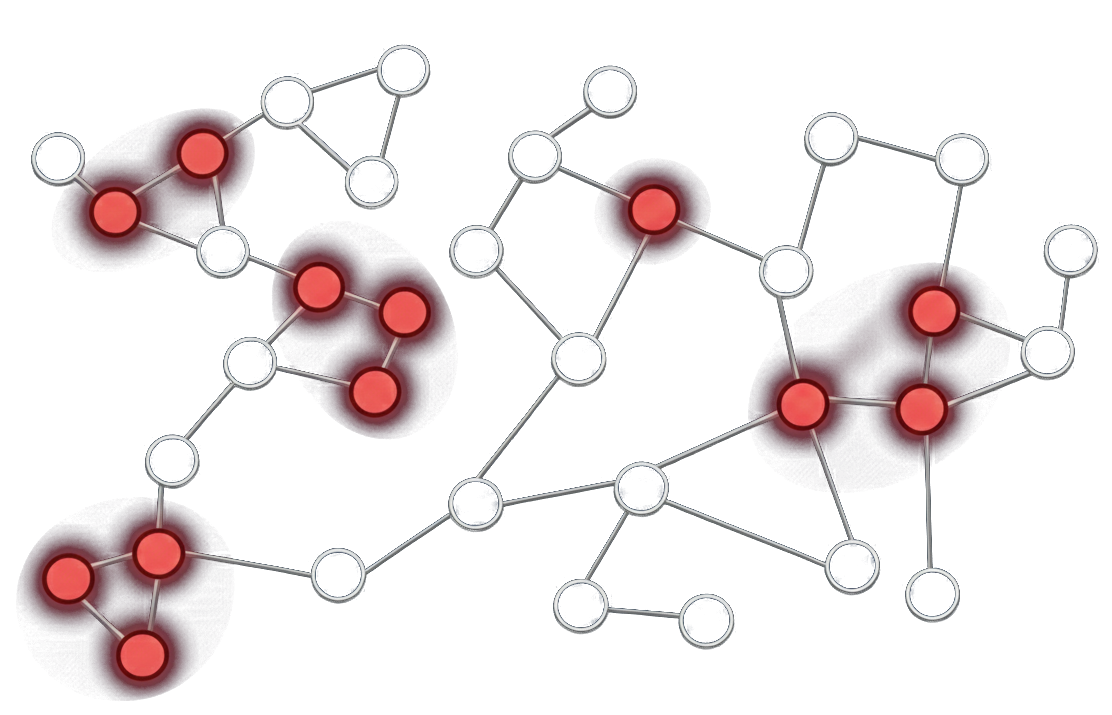
Understanding how genes are regulated requires linking transcriptional programs to underlying chromatin states, yet most single-cell studies profile these modalities separately. We introduce GRIDOT, a framework for reconstructing gene regulatory networks by integrating single-cell RNA-seq and ATAC-seq data without requiring paired measurements. GRIDOT aligns transcriptional and chromatin accessibility profiles to create a pseudo-multiomic representation, enabling the inference of directed regulatory relationships. The method identifies cis-regulatory element–gene and transcription factor–gene interactions and assembles them into regulatory networks at cell-type–specific or population scales. By connecting chromatin regulation to gene expression in a unified framework, GRIDOT facilitates biological interpretation of regulatory mechanisms from unpaired single-cell multiomic data. GRIDOT:
- Aligns distributions across conditions to stabilize downstream causal estimation.
- Infers directed edges with interpretable causal scores.
- Designed for noisy, high-dimensional omics regimes.
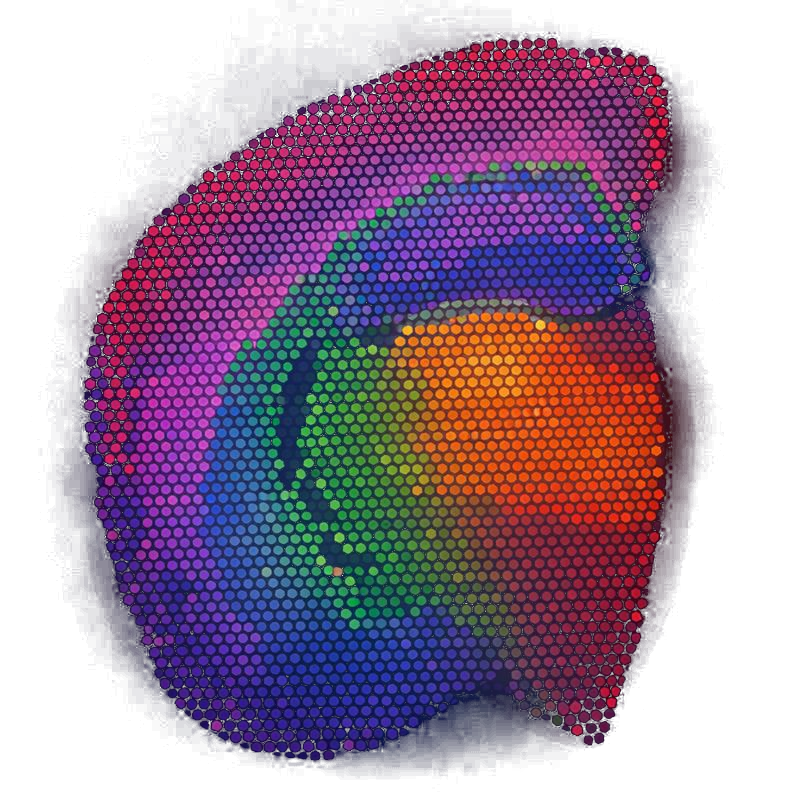
DRIFT
Diffusion-based representation integration for spatial foundation models
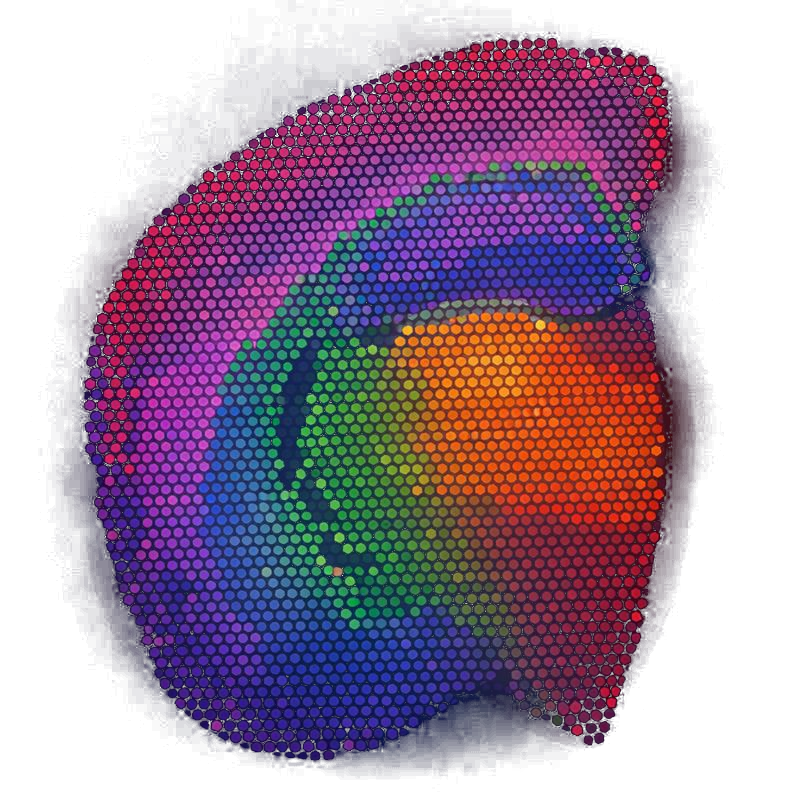
DRIFT
Diffusion-based representation integration for spatial foundation models
DRIFT is a scalable diffusion framework that denoises expression profiles and integrates the spatial topology of ST data into existing pretrained scRNA-seq and ST foundation models without additional retraining. Foundation models that do not explicitly model spatial information benefit from both denoising and spatial integration, while methods that do so leverage DRIFT's denoised output. DRIFT constructs a spatial adjacency graph among tissue spots and applies a heat-kernel diffusion process that propagates gene-expression signals across local neighborhoods while preserving tissue boundaries. This produces spatially coherent yet biologically meaningful representations that can be directly embedded into pretrained foundation models without retraining, making our approach much more computationally scalable and accessible. DRIFT can:
- Enhance neighborhood consistency while preserving biological variation.
- Support robust downstream clustering and spatial pattern recovery.
Odor-associated Learning & Memory
Modeling activity-dependent regulation across odor conditions

Odor-associated Learning & Memory
Modeling activity-dependent regulation across odor conditions
Modeling gene regulatory mechanisms in olfactory cortex and hypothalamus under distinct odor paradigms, focusing on activity-dependent programs and candidate regulators.
- Contrasts condition-specific transcriptional programs.
- Prioritizes candidate regulators for follow-up.
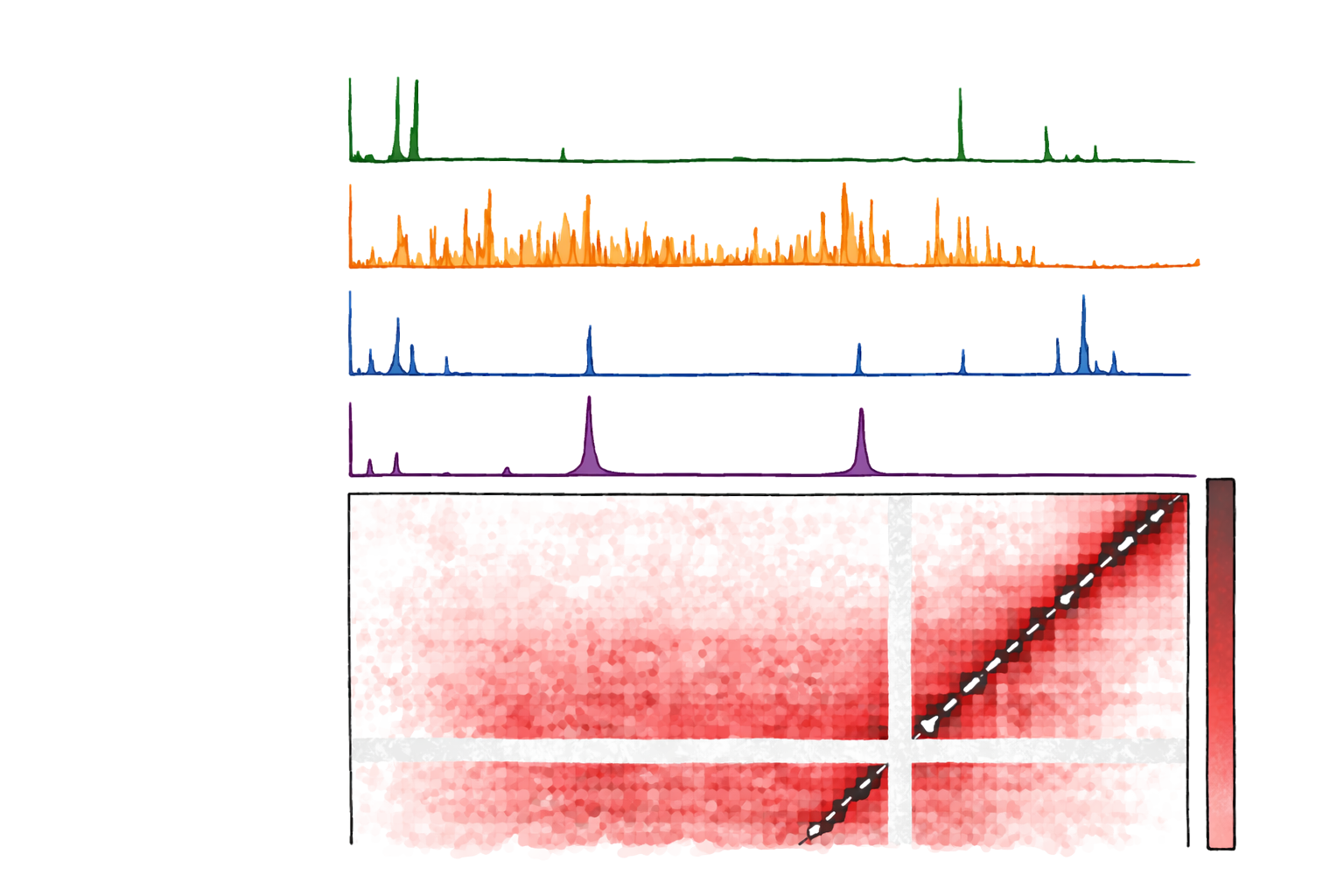
Synergistic TF Control in Drosophila
Multimodal modeling of TF synergy using ChIP-seq and Micro-C

Synergistic TF Control in Drosophila
Multimodal modeling of TF synergy using ChIP-seq and Micro-C
Multimodal modeling of Drosophila transcription factors involved in synapse formation and dosage compensation, integrating sequence, binding, and 3D genome features for interpretable prediction.
- Combines binding profiles with 3D contact structure.
- Targets interpretable TF interactions and combinatorial effects.